 Transcriptional regulation in mammals was simplistically viewed as a process in which soluble protein factors are recruited to promoters, located immediately 5′ of a protein encoding gene, to activate or repress transcription. However, more recent data suggest that the nucleus is an ordered three-dimensional organelle, in which the organization of the genome appears to have implications for orchestrated gene expression. Data suggest that the expression of some genes correlates with their genomic environment. This environment consists of interactions of DNA loci with nuclear structural components such as the nuclear lamina and the nucleoporins and interaction of DNA loci with remote sequences on the same or other chromosomes.
Transcriptional regulation in mammals was simplistically viewed as a process in which soluble protein factors are recruited to promoters, located immediately 5′ of a protein encoding gene, to activate or repress transcription. However, more recent data suggest that the nucleus is an ordered three-dimensional organelle, in which the organization of the genome appears to have implications for orchestrated gene expression. Data suggest that the expression of some genes correlates with their genomic environment. This environment consists of interactions of DNA loci with nuclear structural components such as the nuclear lamina and the nucleoporins and interaction of DNA loci with remote sequences on the same or other chromosomes.
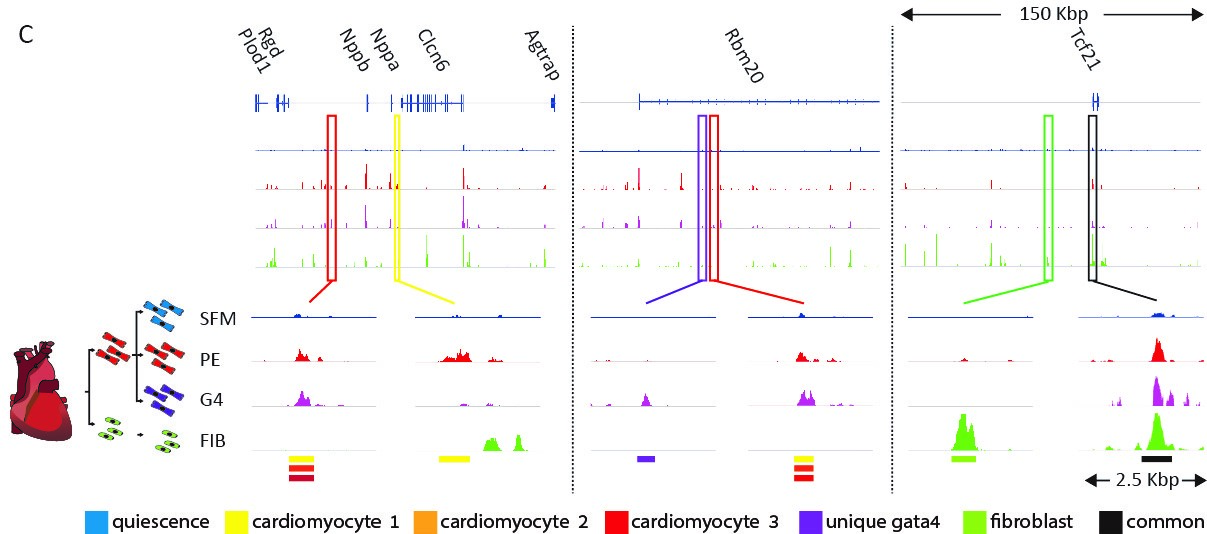
We are trying to understand how cell-type-specific gene expression is controlled by the genome and chromatin in the heart. We performed a genome-wide analysis of regulatory elements (enhancers) in cardiomyocytes and in cardiac fibroblasts using ATAC-seq, H3K27ac ChIP-seq, RNA-seq, and computational transcription factor binding analysis. These analyses enabled us to map the cell-type-specific active enhancers in cardiac fibroblasts and cardiomyocytes and outline the transcription factor families that control them. Our studies showed that in both cell types, distal enhancers, containing concentrated combinatorial clusters of multiple tissue expressed transcription factor recognition motifs, are combinatorically clustered around tissue specific genes.
Paper be Tal Golan-Lagziel (MD/PhD candidate):
Golan-Lagziel T, Lewis YE, Shkedi O, Douvdevany G, Caspi LH, Kehat I. Analysis of rat cardiac myocytes and fibroblasts identifies combinatorial enhancer organization and transcription factor families. J Mol Cell Cardiol. 2018 Mar;116:91-105.
https://www.ncbi.nlm.nih.gov/pubmed/?term=kehat+and+golan-lagziel
https://www.sciencedirect.com/science/article/pii/S0022282818300373?via%3Dihub